MLH1
DNA mismatch repair protein Mlh1 or MutL protein homolog 1 is a protein that in humans is encoded by the MLH1 gene located on chromosome 3. The gene is commonly associated with hereditary nonpolyposis colorectal cancer. Orthologs of human MLH1 have also been studied in other organisms including mouse and the budding yeast Saccharomyces cerevisiae.
Function
[edit]Variants in this gene can cause hereditary nonpolyposis colon cancer (Lynch syndrome). It is a human homolog of the E. coli DNA mismatch repair gene, mutL, which mediates protein-protein interactions during mismatch recognition, strand discrimination, and strand removal. Defects in MLH1 are associated with the microsatellite instability observed in hereditary nonpolyposis colon cancer. Alternatively spliced transcript variants encoding different isoforms have been described, but their full-length natures have not been determined.[4]
Role in DNA mismatch repair
[edit]MLH1 protein is one component of a system of seven DNA mismatch repair proteins that work coordinately in sequential steps to initiate repair of DNA mismatches in humans.[5] Defects in mismatch repair, found in about 13% of colorectal cancers, are much more frequently due to deficiency of MLH1 than deficiencies of other DNA mismatch repair proteins.[6] The seven DNA mismatch repair proteins in humans are MLH1, MLH3, MSH2, MSH3, MSH6, PMS1 and PMS2.[5] In addition, there are Exo1-dependent and Exo1-independent DNA mismatch repair subpathways.[7]
DNA mismatches occur where one base is improperly paired with another base, or where there is a short addition or deletion in one strand of DNA that is not matched in the other strand. Mismatches commonly occur as a result of DNA replication errors or during genetic recombination. Recognizing those mismatches and repairing them is important for cells because failure to do so results in microsatellite instability] and an elevated spontaneous mutation rate (mutator phenotype). Among 20 cancers evaluated, microsatellite instable colon cancer (mismatch repair deficient) had the second highest frequency of mutations (after melanoma).
A heterodimer between MSH2 and MSH6 first recognizes the mismatch, although a heterodimer between MSH2 and MSH3 also can start the process. The formation of the MSH2-MSH6 heterodimer accommodates a second heterodimer of MLH1 and PMS2, although a heterodimer between MLH1 and either PMS3 or MLH3 can substitute for PMS2. This protein complex formed between the 2 sets of heterodimers enables initiation of repair of the mismatch defect.[5]
Other gene products involved in mismatch repair (subsequent to initiation by DNA mismatch repair genes) include DNA polymerase delta, PCNA, RPA, HMGB1, RFC and DNA ligase I, plus histone and chromatin modifying factors.[8][9]
Deficient expression in cancer
[edit]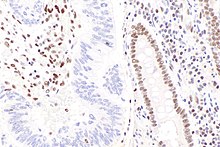
| Cancer type | Frequency of deficiency in cancer | Frequency of deficiency in adjacent field defect |
|---|---|---|
| Stomach | 32%[10][11] | 24–28% |
| Stomach (foveolar type tumors) | 74%[12] | 71% |
| Stomach in high-incidence Kashmir Valley | 73%[13] | 20% |
| Esophageal | 73%[14] | 27% |
| Head and neck squamous cell carcinoma (HNSCC) | 31–33%[15][16] | 20–25% |
| Non-small cell lung cancer (NSCLC) | 69%[17] | 72% |
| Colorectal | 10%[6] |
Epigenetic repression
[edit]Only a minority of sporadic cancers with a DNA repair deficiency have a mutation in a DNA repair gene. However, a majority of sporadic cancers with a DNA repair deficiency do have one or more epigenetic alterations that reduce or silence DNA repair gene expression.[18] In the table above, the majority of deficiencies of MLH1 were due to methylation of the promoter region of the MLH1 gene. Another epigenetic mechanism reducing MLH1 expression is over-expression of miR-155.[19] MiR-155 targets MLH1 and MSH2 and an inverse correlation between the expression of miR-155 and the expression of MLH1 or MSH2 proteins was found in human colorectal cancer.[19]
Deficiency in field defects
[edit]A field defect is an area or "field" of epithelium that has been preconditioned by epigenetic changes and/or mutations so as to predispose it towards development of cancer. As pointed out by Rubin, "The vast majority of studies in cancer research has been done on well-defined tumors in vivo, or on discrete neoplastic foci in vitro.[20] Yet there is evidence that more than 80% of the somatic mutations found in mutator phenotype human colorectal tumors occur before the onset of terminal clonal expansion."[21] Similarly, Vogelstein et al.[22] point out that more than half of somatic mutations identified in tumors occurred in a pre-neoplastic phase (in a field defect), during growth of apparently normal cells.
In the Table above, MLH1 deficiencies were noted in the field defects (histologically normal tissues) surrounding most of the cancers. If MLH1 is epigenetically reduced or silenced, it would not likely confer a selective advantage upon a stem cell. However, reduced or absent expression of MLH1 would cause increased rates of mutation, and one or more of the mutated genes may provide the cell with a selective advantage. The expression-deficient MLH1 gene could then be carried along as a selectively neutral or only slightly deleterious passenger (hitch-hiker) gene when the mutated stem cell generates an expanded clone. The continued presence of a clone with an epigenetically repressed MLH1 would continue to generate further mutations, some of which could produce a tumor.
Repression in coordination with other DNA repair genes
[edit]In a cancer, multiple DNA repair genes are often found to be simultaneously repressed.[18] In one example, involving MLH1, Jiang et al.[23] conducted a study where they evaluated the mRNA expression of 27 DNA repair genes in 40 astrocytomas compared to normal brain tissues from non-astrocytoma individuals. Among the 27 DNA repair genes evaluated, 13 DNA repair genes, MLH1, MLH3, MGMT, NTHL1, OGG1, SMUG1, ERCC1, ERCC2, ERCC3, ERCC4, RAD50, XRCC4 and XRCC5 were all significantly down-regulated in all three grades (II, III and IV) of astrocytomas. The repression of these 13 genes in lower grade as well as in higher grade astrocytomas suggested that they may be important in early as well as in later stages of astrocytoma. In another example, Kitajima et al.[24] found that immunoreactivity for MLH1 and MGMT expression was closely correlated in 135 specimens of gastric cancer and loss of MLH1 and MGMTappeared to be synchronously accelerated during tumor progression.
Deficient expression of multiple DNA repair genes are often found in cancers,[18] and may contribute to the thousands of mutations usually found in cancers (see Mutation frequencies in cancers).
Meiosis
[edit]In addition to its role in DNA mismatch repair, MLH1 protein is also involved in meiotic crossing over.[25] MLH1 forms a heterodimer with MLH3 that appears to be necessary for oocytes to progress through metaphase II of meiosis.[26] Female and male MLH1(-/-) mutant mice are infertile, and sterility is associated with a reduced level of chiasmata.[25][27] During spermatogenesis in MLH1(-/-) mutant mice chromosomes often separate prematurely and there is frequent arrest in the first division of meiosis.[25] In humans, a common variant of the MLH1 gene is associated with increased risk of sperm damage and male infertility.[28]

MLH1 protein appears to localize to sites of crossing over in meiotic chromosomes.[25] Recombination during meiosis is often initiated by a DNA double-strand break (DSB) as illustrated in the accompanying diagram. During recombination, sections of DNA at the 5' ends of the break are cut away in a process called resection. In the strand invasion step that follows, an overhanging 3' end of the broken DNA molecule then "invades" the DNA of an homologous chromosome that is not broken forming a displacement loop (D-loop). After strand invasion, the further sequence of events may follow either of two main pathways leading to a crossover (CO) or a non-crossover (NCO) recombinant (see Genetic recombination). The pathway leading to a CO involves a double Holliday junction (DHJ) intermediate. Holliday junctions need to be resolved for CO recombination to be completed.
In the budding yeast Saccharomyces cerevisiae, as in the mouse, MLH1 forms a heterodimer with MLH3. Meiotic CO requires resolution of Holliday junctions through actions of the MLH1-MLH3 heterodimer. The MLH1-MLH3 heterodimer is an endonuclease that makes single-strand breaks in supercoiled double-stranded DNA.[29][30] MLH1-MLH3 binds specifically to Holliday junctions and may act as part of a larger complex to process Holliday junctions during meiosis.[29] MLH1-MLH3 heterodimer (MutL gamma) together with EXO1 and Sgs1 (ortholog of Bloom syndrome helicase) define a joint molecule resolution pathway that produces the majority of crossovers in budding yeast and, by inference, in mammals.[31]
Clinical significance
[edit]It can also be associated with Turcot syndrome.[32]
Interactions
[edit]MLH1 has been shown to interact with:
See also
[edit]References
[edit]- ^ a b c GRCm38: Ensembl release 89: ENSMUSG00000032498 – Ensembl, May 2017
- ^ "Human PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Mouse PubMed Reference:". National Center for Biotechnology Information, U.S. National Library of Medicine.
- ^ "Entrez Gene: MLH1 mutL homolog 1, colon cancer, nonpolyposis type 2 (E. coli)".
- ^ a b c Pal T, Permuth-Wey J, Sellers TA (2008). "A review of the clinical relevance of mismatch-repair deficiency in ovarian cancer". Cancer. 113 (4): 733–42. doi:10.1002/cncr.23601. PMC 2644411. PMID 18543306.
- ^ a b Truninger K, Menigatti M, Luz J, Russell A, Haider R, Gebbers JO, Bannwart F, Yurtsever H, Neuweiler J, Riehle HM, Cattaruzza MS, Heinimann K, Schär P, Jiricny J, Marra G (2005). "Immunohistochemical analysis reveals high frequency of PMS2 defects in colorectal cancer". Gastroenterology. 128 (5): 1160–71. doi:10.1053/j.gastro.2005.01.056. PMID 15887099.
- ^ Goellner EM, Putnam CD, Kolodner RD (2015). "Exonuclease 1-dependent and independent mismatch repair". DNA Repair (Amst.). 32: 24–32. doi:10.1016/j.dnarep.2015.04.010. PMC 4522362. PMID 25956862.
- ^ Li GM (2008). "Mechanisms and functions of DNA mismatch repair". Cell Res. 18 (1): 85–98. doi:10.1038/cr.2007.115. PMID 18157157.
- ^ Li GM (2014). "New insights and challenges in mismatch repair: getting over the chromatin hurdle". DNA Repair (Amst.). 19: 48–54. doi:10.1016/j.dnarep.2014.03.027. PMC 4127414. PMID 24767944.
- ^ Kupčinskaitė-Noreikienė R, Skiecevičienė J, Jonaitis L, Ugenskienė R, Kupčinskas J, Markelis R, Baltrėnas V, Sakavičius L, Semakina I, Grižas S, Juozaitytė E (2013). "CpG island methylation of the MLH1, MGMT, DAPK, and CASP8 genes in cancerous and adjacent noncancerous stomach tissues". Medicina (Kaunas). 49 (8): 361–6. PMID 24509146.
- ^ Waki T, Tamura G, Tsuchiya T, Sato K, Nishizuka S, Motoyama T (2002). "Promoter methylation status of E-cadherin, hMLH1, and p16 genes in nonneoplastic gastric epithelia". Am. J. Pathol. 161 (2): 399–403. doi:10.1016/S0002-9440(10)64195-8. PMC 1850716. PMID 12163364.
- ^ Endoh Y, Tamura G, Ajioka Y, Watanabe H, Motoyama T (2000). "Frequent hypermethylation of the hMLH1 gene promoter in differentiated-type tumors of the stomach with the gastric foveolar phenotype". Am. J. Pathol. 157 (3): 717–22. doi:10.1016/S0002-9440(10)64584-1. PMC 1949419. PMID 10980110.
- ^ Wani M, Afroze D, Makhdoomi M, Hamid I, Wani B, Bhat G, Wani R, Wani K (2012). "Promoter methylation status of DNA repair gene (hMLH1) in gastric carcinoma patients of the Kashmir valley". Asian Pac. J. Cancer Prev. 13 (8): 4177–81. doi:10.7314/apjcp.2012.13.8.4177. PMID 23098428.
- ^ Chang Z, Zhang W, Chang Z, Song M, Qin Y, Chang F, Guo H, Wei Q (2015). "Expression characteristics of FHIT, p53, BRCA2 and MLH1 in families with a history of oesophageal cancer in a region with a high incidence of oesophageal cancer". Oncol Lett. 9 (1): 430–436. doi:10.3892/ol.2014.2682. PMC 4246613. PMID 25436004.
- ^ Tawfik HM, El-Maqsoud NM, Hak BH, El-Sherbiny YM (2011). "Head and neck squamous cell carcinoma: mismatch repair immunohistochemistry and promoter hypermethylation of hMLH1 gene". Am J Otolaryngol. 32 (6): 528–36. doi:10.1016/j.amjoto.2010.11.005. PMID 21353335.
- ^ Zuo C, Zhang H, Spencer HJ, Vural E, Suen JY, Schichman SA, Smoller BR, Kokoska MS, Fan CY (2009). "Increased microsatellite instability and epigenetic inactivation of the hMLH1 gene in head and neck squamous cell carcinoma". Otolaryngol Head Neck Surg. 141 (4): 484–90. doi:10.1016/j.otohns.2009.07.007. PMID 19786217. S2CID 8357370.
- ^ Safar AM, Spencer H, Su X, Coffey M, Cooney CA, Ratnasinghe LD, Hutchins LF, Fan CY (2005). "Methylation profiling of archived non-small cell lung cancer: a promising prognostic system". Clin. Cancer Res. 11 (12): 4400–5. doi:10.1158/1078-0432.CCR-04-2378. PMID 15958624.
- ^ a b c Bernstein C, Bernstein H (2015). "Epigenetic reduction of DNA repair in progression to gastrointestinal cancer". World J Gastrointest Oncol. 7 (5): 30–46. doi:10.4251/wjgo.v7.i5.30. PMC 4434036. PMID 25987950.
- ^ a b Valeri N, Gasparini P, Fabbri M, Braconi C, Veronese A, Lovat F, Adair B, Vannini I, Fanini F, Bottoni A, Costinean S, Sandhu SK, Nuovo GJ, Alder H, Gafa R, Calore F, Ferracin M, Lanza G, Volinia S, Negrini M, McIlhatton MA, Amadori D, Fishel R, Croce CM (2010). "Modulation of mismatch repair and genomic stability by miR-155". Proc. Natl. Acad. Sci. U.S.A. 107 (15): 6982–7. Bibcode:2010PNAS..107.6982V. doi:10.1073/pnas.1002472107. PMC 2872463. PMID 20351277.
- ^ Rubin H (March 2011). "Fields and field cancerization: the preneoplastic origins of cancer: asymptomatic hyperplastic fields are precursors of neoplasia, and their progression to tumors can be tracked by saturation density in culture". BioEssays. 33 (3): 224–31. doi:10.1002/bies.201000067. PMID 21254148. S2CID 44981539.
- ^ Tsao JL, Yatabe Y, Salovaara R, Järvinen HJ, Mecklin JP, Aaltonen LA, Tavaré S, Shibata D (February 2000). "Genetic reconstruction of individual colorectal tumor histories". Proc. Natl. Acad. Sci. U.S.A. 97 (3): 1236–41. Bibcode:2000PNAS...97.1236T. doi:10.1073/pnas.97.3.1236. PMC 15581. PMID 10655514.
- ^ Vogelstein B, Papadopoulos N, Velculescu VE, Zhou S, Diaz LA, Kinzler KW (March 2013). "Cancer genome landscapes". Science. 339 (6127): 1546–58. Bibcode:2013Sci...339.1546V. doi:10.1126/science.1235122. PMC 3749880. PMID 23539594.
- ^ Jiang Z, Hu J, Li X, Jiang Y, Zhou W, Lu D (2006). "Expression analyses of 27 DNA repair genes in astrocytoma by TaqMan low-density array". Neurosci. Lett. 409 (2): 112–7. doi:10.1016/j.neulet.2006.09.038. PMID 17034947. S2CID 54278905.
- ^ Kitajima Y, Miyazaki K, Matsukura S, Tanaka M, Sekiguchi M (2003). "Loss of expression of DNA repair enzymes MGMT, hMLH1, and hMSH2 during tumor progression in gastric cancer". Gastric Cancer. 6 (2): 86–95. doi:10.1007/s10120-003-0213-z. PMID 12861399.
- ^ a b c d Baker SM, Plug AW, Prolla TA, Bronner CE, Harris AC, Yao X, Christie DM, Monell C, Arnheim N, Bradley A, Ashley T, Liskay RM (1996). "Involvement of mouse Mlh1 in DNA mismatch repair and meiotic crossing over". Nat. Genet. 13 (3): 336–42. doi:10.1038/ng0796-336. PMID 8673133. S2CID 37096830.
- ^ Kan R, Sun X, Kolas NK, Avdievich E, Kneitz B, Edelmann W, Cohen PE (2008). "Comparative analysis of meiotic progression in female mice bearing mutations in genes of the DNA mismatch repair pathway". Biol. Reprod. 78 (3): 462–71. doi:10.1095/biolreprod.107.065771. PMID 18057311.
- ^ Wei K, Kucherlapati R, Edelmann W (2002). "Mouse models for human DNA mismatch-repair gene defects". Trends Mol Med. 8 (7): 346–53. doi:10.1016/s1471-4914(02)02359-6. PMID 12114115.
- ^ Ji G, Long Y, Zhou Y, Huang C, Gu A, Wang X (2012). "Common variants in mismatch repair genes associated with increased risk of sperm DNA damage and male infertility". BMC Med. 10: 49. doi:10.1186/1741-7015-10-49. PMC 3378460. PMID 22594646.
- ^ a b Ranjha L, Anand R, Cejka P (2014). "The Saccharomyces cerevisiae Mlh1-Mlh3 heterodimer is an endonuclease that preferentially binds to Holliday junctions". J. Biol. Chem. 289 (9): 5674–86. doi:10.1074/jbc.M113.533810. PMC 3937642. PMID 24443562.
- ^ Rogacheva MV, Manhart CM, Chen C, Guarne A, Surtees J, Alani E (2014). "Mlh1-Mlh3, a meiotic crossover and DNA mismatch repair factor, is a Msh2-Msh3-stimulated endonuclease". J. Biol. Chem. 289 (9): 5664–73. doi:10.1074/jbc.M113.534644. PMC 3937641. PMID 24403070.
- ^ Zakharyevich K, Tang S, Ma Y, Hunter N (2012). "Delineation of joint molecule resolution pathways in meiosis identifies a crossover-specific resolvase". Cell. 149 (2): 334–47. doi:10.1016/j.cell.2012.03.023. PMC 3377385. PMID 22500800.
- ^ Lebrun C, Olschwang S, Jeannin S, Vandenbos F, Sobol H, Frenay M (2007). "Turcot syndrome confirmed with molecular analysis". Eur. J. Neurol. 14 (4): 470–2. doi:10.1111/j.1468-1331.2006.01669.x. PMID 17389002. S2CID 21591979.
- ^ Wang Y, Cortez D, Yazdi P, Neff N, Elledge SJ, Qin J (April 2000). "BASC, a super complex of BRCA1-associated proteins involved in the recognition and repair of aberrant DNA structures". Genes Dev. 14 (8): 927–39. doi:10.1101/gad.14.8.927. PMC 316544. PMID 10783165.
- ^ Langland G, Kordich J, Creaney J, Goss KH, Lillard-Wetherell K, Bebenek K, Kunkel TA, Groden J (August 2001). "The Bloom's syndrome protein (BLM) interacts with MLH1 but is not required for DNA mismatch repair". J. Biol. Chem. 276 (32): 30031–5. doi:10.1074/jbc.M009664200. PMID 11325959.
- ^ Freire R, d'Adda Di Fagagna F, Wu L, Pedrazzi G, Stagljar I, Hickson ID, Jackson SP (August 2001). "Cleavage of the Bloom's syndrome gene product during apoptosis by caspase-3 results in an impaired interaction with topoisomerase IIIalpha". Nucleic Acids Res. 29 (15): 3172–80. doi:10.1093/nar/29.15.3172. PMC 55826. PMID 11470874.
- ^ Pedrazzi G, Perrera C, Blaser H, Kuster P, Marra G, Davies SL, Ryu GH, Freire R, Hickson ID, Jiricny J, Stagljar I (November 2001). "Direct association of Bloom's syndrome gene product with the human mismatch repair protein MLH1". Nucleic Acids Res. 29 (21): 4378–86. doi:10.1093/nar/29.21.4378. PMC 60193. PMID 11691925.
- ^ Schmutte C, Sadoff MM, Shim KS, Acharya S, Fishel R (August 2001). "The interaction of DNA mismatch repair proteins with human exonuclease I". J. Biol. Chem. 276 (35): 33011–8. doi:10.1074/jbc.M102670200. PMID 11427529.
- ^ Bellacosa A, Cicchillitti L, Schepis F, Riccio A, Yeung AT, Matsumoto Y, Golemis EA, Genuardi M, Neri G (March 1999). "MED1, a novel human methyl-CpG-binding endonuclease, interacts with DNA mismatch repair protein MLH1". Proc. Natl. Acad. Sci. U.S.A. 96 (7): 3969–74. Bibcode:1999PNAS...96.3969B. doi:10.1073/pnas.96.7.3969. PMC 22404. PMID 10097147.
- ^ Santucci-Darmanin S, Walpita D, Lespinasse F, Desnuelle C, Ashley T, Paquis-Flucklinger V (August 2000). "MSH4 acts in conjunction with MLH1 during mammalian meiosis". FASEB J. 14 (11): 1539–47. doi:10.1096/fj.14.11.1539. PMID 10928988.
- ^ a b Mac Partlin M, Homer E, Robinson H, McCormick CJ, Crouch DH, Durant ST, Matheson EC, Hall AG, Gillespie DA, Brown R (February 2003). "Interactions of the DNA mismatch repair proteins MLH1 and MSH2 with c-MYC and MAX". Oncogene. 22 (6): 819–25. doi:10.1038/sj.onc.1206252. PMID 12584560.
- ^ Kondo E, Horii A, Fukushige S (April 2001). "The interacting domains of three MutL heterodimers in man: hMLH1 interacts with 36 homologous amino acid residues within hMLH3, hPMS1 and hPMS2". Nucleic Acids Res. 29 (8): 1695–702. doi:10.1093/nar/29.8.1695. PMC 31313. PMID 11292842.
- ^ Guerrette S, Acharya S, Fishel R (March 1999). "The interaction of the human MutL homologues in hereditary nonpolyposis colon cancer". J. Biol. Chem. 274 (10): 6336–41. doi:10.1074/jbc.274.10.6336. PMID 10037723.
Further reading
[edit]- Paraf F, Sasseville D, Watters AK, Narod S, Ginsburg O, Shibata H, Jothy S (1995). "Clinicopathological relevance of the association between gastrointestinal and sebaceous neoplasms: the Muir-Torre syndrome". Hum. Pathol. 26 (4): 422–7. doi:10.1016/0046-8177(95)90144-2. PMID 7705822.
- Kolodner RD (1996). "Mismatch repair: mechanisms and relationship to cancer susceptibility". Trends Biochem. Sci. 20 (10): 397–401. doi:10.1016/S0968-0004(00)89087-8. PMID 8533151.
- Peltomäki P, de la Chapelle A (1997). Mutations predisposing to hereditary nonpolyposis colorectal cancer. Advances in Cancer Research. Vol. 71. pp. 93–119. doi:10.1016/S0065-230X(08)60097-4. ISBN 9780120066711. PMID 9111864.
{{cite book}}:|journal=ignored (help) - Papadopoulos N, Lindblom A (1997). "Molecular basis of HNPCC: mutations of MMR genes". Hum. Mutat. 10 (2): 89–99. doi:10.1002/(SICI)1098-1004(1997)10:2<89::AID-HUMU1>3.0.CO;2-H. PMID 9259192. S2CID 6799575.
- Kauh J, Umbreit J (2004). "Colorectal cancer prevention". Current Problems in Cancer. 28 (5): 240–64. doi:10.1016/j.currproblcancer.2004.05.004. PMID 15375803.
- Warusavitarne J, Schnitzler M (2007). "The role of chemotherapy in microsatellite unstable (MSI-H) colorectal cancer". International Journal of Colorectal Disease. 22 (7): 739–48. doi:10.1007/s00384-006-0228-0. PMID 17109103. S2CID 6460105.
- Niv Y (2007). "Microsatellite instability and MLH1 promoter hypermethylation in colorectal cancer". World J. Gastroenterol. 13 (12): 1767–9. doi:10.3748/wjg.v13.i12.1767. PMC 4149951. PMID 17465465.
External links
[edit]- Information on HNPCC from the National Institute of Health
- GeneReviews/NCBI/NIH/UW entry on Lynch syndrome
- MLH1+protein,+human at the U.S. National Library of Medicine Medical Subject Headings (MeSH)
- Overview of all the structural information available in the PDB for UniProt: P40692 (DNA mismatch repair protein Mlh1) at the PDBe-KB.


 French
French Deutsch
Deutsch

